Software / 2026
Molecule Detection, Annotation and Analysis (MolDetA)
PyQt desktop application for synchronized STM datasets: registration, molecule detection, manual annotation, grouping, acquisition-aware analysis and data export.

Overview
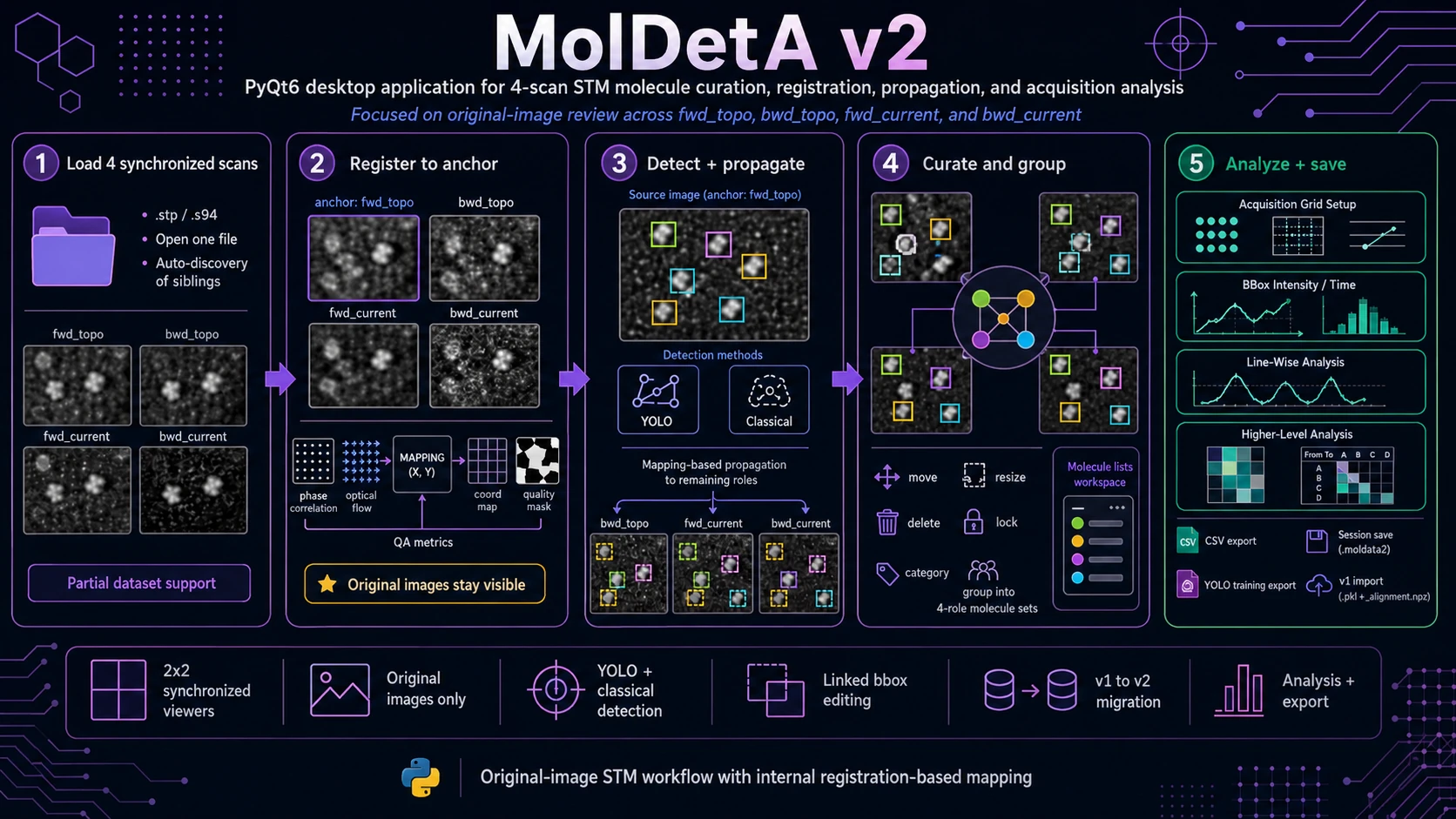
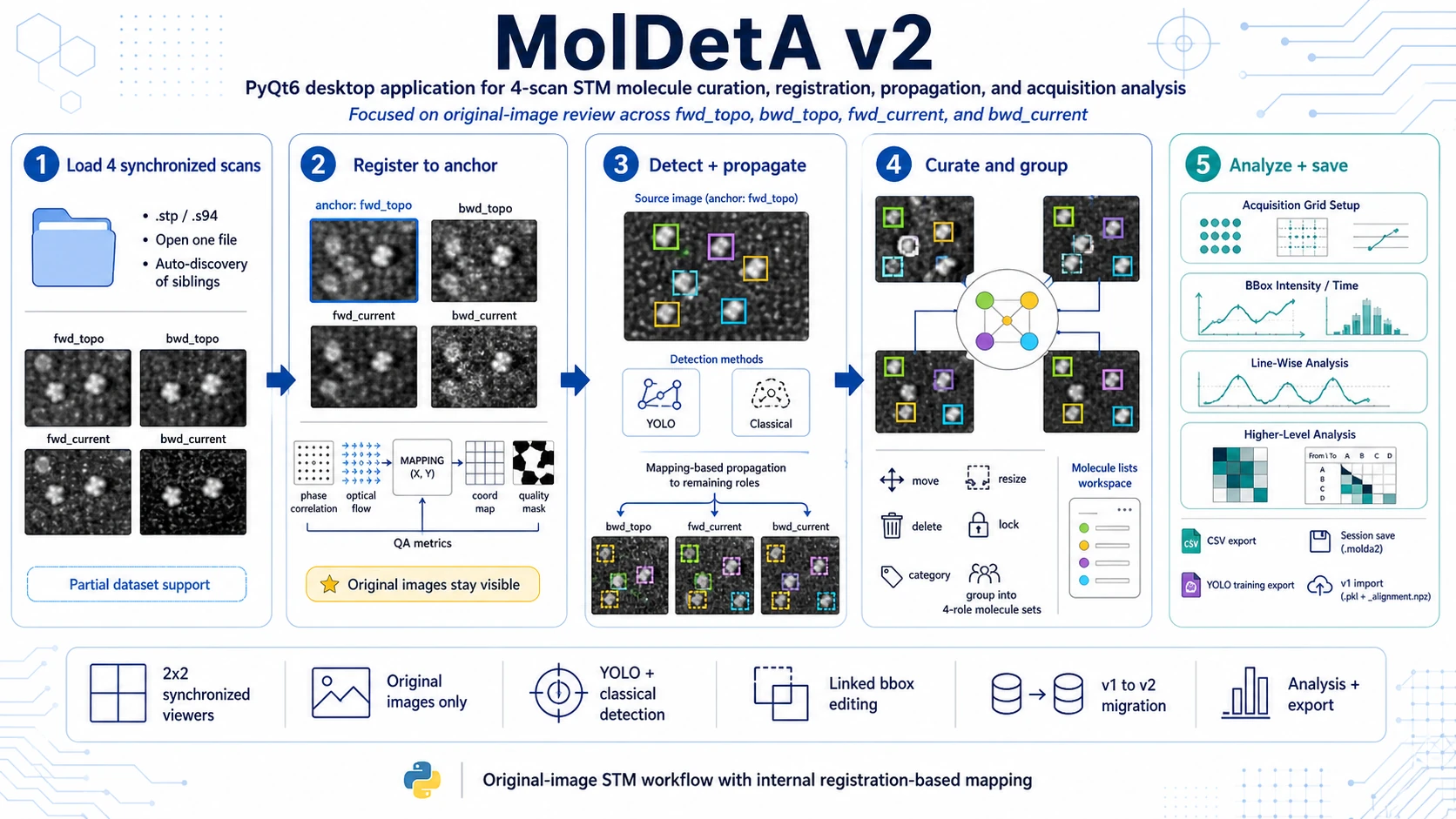
Molecule Detection, Annotation and Analysis (MolDetA) v2 is a PyQt desktop application for molecule-level curation and analysis of synchronized STM datasets. It is designed for experiments where the same area is acquired as a four-scan family:
fwd_topo- forward topography,bwd_topo- backward topography,fwd_current- forward current,bwd_current- backward current.
The central design rule is that all viewers show the original, non-warped STM images. Registration is used internally only as a coordinate-mapping layer for propagating molecule positions, checking consistency and running downstream analysis. This keeps visual review grounded in the measured data while still allowing annotations to stay synchronized across scan direction and signal channel.
What MolDetA v2 Does
MolDetA v2 helps the user move from raw STM scans to curated molecule sets and analysis-ready tables. The software combines four-scan data loading, registration, detection, manual annotation, grouping, category review, acquisition-grid analysis and export in one workflow.
At a high level, MolDetA v2 provides:
- single-file STM loading with sibling discovery for the four scan roles,
2×2synchronized viewers for simultaneous inspection of forward/backward topography and current scans,- internal registration and coordinate mapping based on rigid pre-alignment and optical flow,
- source-scan molecule detection using YOLO or classical image-processing pipelines,
- mapping-based bbox propagation from the selected source scan to the remaining scan roles,
- manual bbox curation with move, resize, delete, lock, category and linked-edit workflows,
- four-role molecule grouping for building consistent molecule records across scan roles,
- detached molecule-list workspace for review on a second monitor,
- numeric category assignment and filtering for dense-scene review,
- acquisition-grid, timing, intensity and line-wise analysis,
- higher-level analysis including table previews, Hopkins charts, conditional distributions and group-transition views,
- native
.molda2session save/load and import of legacy MolDetA v1 sessions.
Why Original-Image Review Matters
STM scans are sequentially acquired and can differ between scan direction, topography/current channel and tunneling conditions. A visually warped display can make curation harder because the user no longer sees exactly what was measured.
MolDetA v2 avoids this problem:
- the four viewers always show the original STM frames,
- registration outputs are stored as internal mapping artifacts,
- propagated bboxes are overlays computed from the mapping,
- manual corrections remain explicit and are never hidden inside a display transform.
This workflow is suitable for careful molecule-by-molecule review, especially when topography and current contrast differ or when forward/backward scans contain small positional shifts.
Typical Workflow
1. Open One STM File
Use File → Open one STM file… and select one file from a scan family. MolDetA v2 attempts to discover the sibling files that correspond to:
fwd_topo,bwd_topo,fwd_current,bwd_current.
Currently, the v2 editing/runtime workflow is intended for a complete four-image dataset. If a required scan role is missing, the application reports the exact missing role or roles so the dataset can be corrected before analysis continues.
2. Review Registration Quality
Use Tools → Registration Sandbox (4 scans)… to inspect registration before trusting propagated geometry. The sandbox supports anchor-based registration, with fwd_topo as the default anchor. It exposes diagnostics such as shift, optical-flow behavior, residuals, masks and mapping-quality indicators.
The registration layer is used to map coordinates between scan roles, not to replace the displayed images.
3. Detect Molecules on a Source Scan
Choose an active source image and run a detector. MolDetA v2 supports two detection families:
- YOLO detection for model-based molecule proposals,
- classical detection for algorithmic detection using image-processing steps such as preprocessing, local maxima, Difference of Gaussians and Laplacian of Gaussian style workflows.
Detections are converted into canonical molecule/bbox objects. The detected centers are then propagated to the remaining roles through the internal coordinate mapping.
4. Curate Bboxes Manually
After detection and propagation, review all bboxes directly on the original STM images. The user can:
- add a bbox manually,
- move or resize a bbox on one scan role,
- move or resize linked bboxes together,
- delete the selected role geometry,
- lock objects to prevent accidental edits,
- assign or clear numeric categories,
- use filters and dimming modes to focus on selected molecule classes.
Manual geometry has priority over propagated or detected geometry. In practice, the source hierarchy is:
manual > propagated > detected5. Group Molecules Across Four Roles
MolDetA v2 represents molecule identity through grouped role-specific bboxes. A group can contain one bbox per scan role. Grouping mode is used to create or edit these relationships across:
- forward topography,
- backward topography,
- forward current,
- backward current.
This allows a single molecule to be followed consistently across scan direction and imaging channel while still preserving the original per-role geometry.
6. Save the Session
Use File → Save Session v2… to save the complete project as a native .molda2 session. A v2 session preserves dataset references, registration artifacts, bbox geometry, grouping state, categories, visual-review state and acquisition-analysis settings.
Use File → Load Session v2… to continue the workflow later.
7. Run Acquisition and Downstream Analysis
When molecule curation is ready, use the analysis tools:
- Acquisition Grid Setup - configure restored/native acquisition grid, orientation, flip settings, offsets, scan-speed context and forward/backward anchor-line synchronization.
- BBox Intensity / Time Analysis - extract molecule-level intensity and timing summaries for single-scan and pairwise views.
- Line-Wise Analysis - extract line-level observations for molecules intersecting acquisition lines or grid cells.
- Higher-Level Analysis - inspect cached BBox/Line-Wise results through tables, Hopkins charts, conditional distributions and group-assignment/transition views.
Analysis CSV exports are available from the BBox and Line-Wise analysis tools.
Main Use Cases
Four-Scan Molecule Curation
Use MolDetA v2 to curate molecule positions across forward/backward and topography/current STM scans while keeping all images visible in their original measured form.
Cross-Channel Molecule Mapping
Detect molecules on the scan role with the clearest contrast, then propagate their bbox centers to the other roles using registration-based mapping. This is useful when the molecule is easier to detect in one channel, but downstream analysis requires comparison across all four scans.
Manual Correction After Automated Detection
Use YOLO or classical detection as a fast first pass, then correct individual bboxes manually. Manual edits are tracked explicitly and take precedence over automatically propagated geometry.
Dense-Scene Review and Categorization
Use numeric categories, filters, dimming, lock states and the molecule-list workspace to review crowded molecule fields. This supports classification, reassignment and quality-control workflows.
Forward/Backward Consistency Analysis
Group molecule bboxes across forward and backward scans to compare geometry, intensity, timing and line-wise observations between scan directions.
Acquisition-Timing and Line-Wise Analysis
Configure an acquisition grid and use BBox or Line-Wise analysis to connect molecule positions with scan timing, acquisition lines and intensity behavior.
Legacy Project Migration
Import historical MolDetA v1 pair-based sessions and continue the project in the v2 four-role model. Legacy sessions are treated as migration inputs; MolDetA v2 is the primary workflow for current work.
Detector Dataset Preparation
Export curated molecule annotations in a YOLO-style images/labels layout for training or improving future detection models.
Inputs and Outputs
Supported STM Inputs
.stp,.s94.
Session Formats
- native MolDetA v2 session:
.molda2, - legacy MolDetA v1 import:
.pklwith companion_alignment.npz.
Export Paths
- YOLO-style training data export from the current curation state,
- CSV export from BBox Intensity / Time Analysis,
- CSV export from Line-Wise Analysis,
- geometric-validation CSV/JSON/Markdown artifacts from the validation script.
Installation and Launch
Install the base dependencies:
pip install -r requirements.txtRecommended optional packages for image-processing workflows:
pip install scikit-image bm3dFor YOLO detection workflows, also install Ultralytics and a compatible PyTorch build:
pip install ultralytics torchRun MolDetA v2:
python -m moldeta_v2.mainMolDetA v2 is developed as a desktop PyQt application. Python 3.10+ is recommended. CPU workflows are supported; an NVIDIA GPU is optional for accelerating YOLO detection.
Relationship to MolDetA v1
The repository also contains the legacy moldeta application. It is useful for historical pair-based workflows and for validating the migration path, but MolDetA v2 is the current version and the target for present testing.
Legacy MolDetA v1 projects can be imported into MolDetA v2 for continued four-scan review and analysis.
Relationship to LFA
MolDetA v2 and LFA address different STM-analysis problems:
- LFA focuses on reciprocal-space and lattice metrology: FFT analysis, affine calibration, substrate/adsorbate lattice parameters, superstructure periods and uncertainty-bearing lattice metrics.
- MolDetA v2 focuses on molecule-level annotation and analysis across synchronized STM scans: detection, bbox propagation, manual curation, grouping, timing/intensity analysis and export.
Together they cover complementary parts of a broader STM/EC-STM analysis workflow.
Current Status
MolDetA v2 is in the testing and stabilization stage. The core implementation contract is functionally complete: loading, registration, detection, propagation, manual editing, grouping, session persistence, v1 migration, acquisition analysis, exports and higher-level review tools are implemented.
The remaining work is focused on validation, regression testing and practical verification on target datasets rather than adding new feature areas.