Software / 2026
NanoTrack
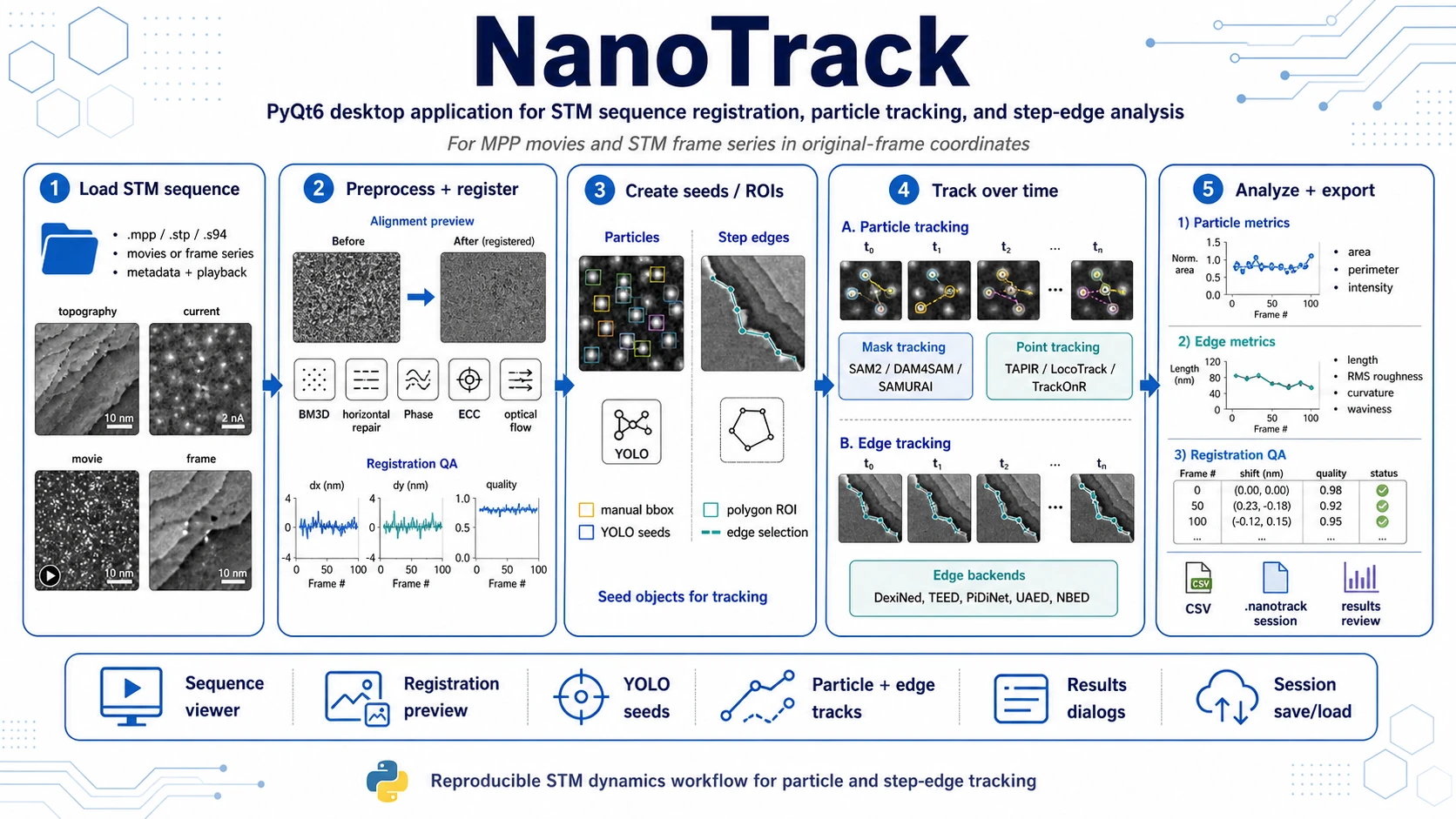
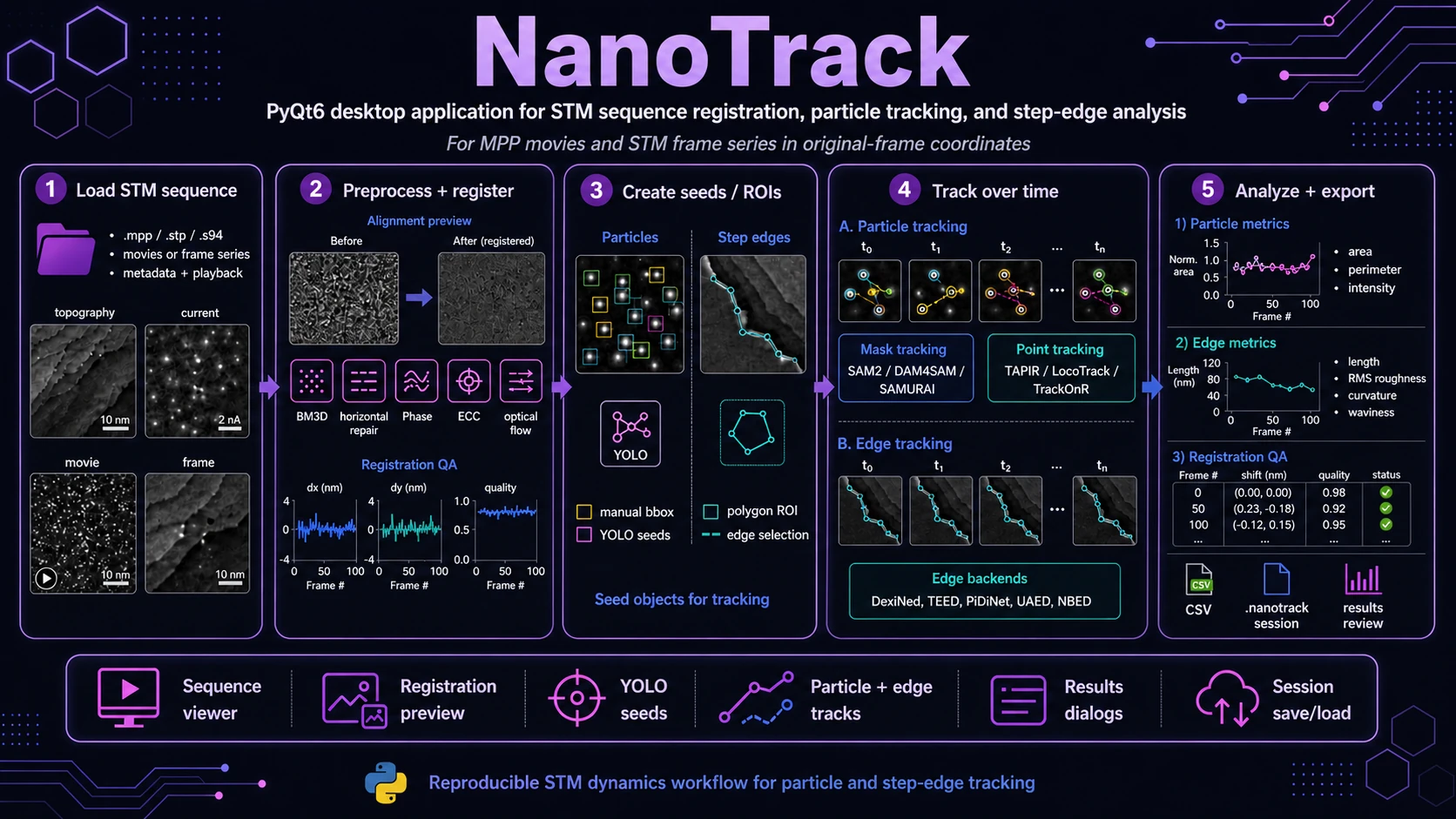
PyQt6 desktop application for time-resolved STM sequence analysis: frame registration, particle and step-edge seeds, tracking, manual review and metric export.
Overview
NanoTrack is a PyQt6 desktop application for reviewing STM frame sequences, registering frames, creating particle and step-edge seeds, tracking them over time and exporting quantitative results.
It is designed for STM movies and frame series where the relevant information is not only the shape of one frame, but also how particles, defects, adsorbates or step edges evolve during the scan sequence.
NanoTrack is part of the same repository ecosystem as NaParA, but it is presented as a separate application. NaParA focuses on nanoparticle detection and per-image quantification, while NanoTrack focuses on time-resolved STM sequence workflows.
What NanoTrack Does
NanoTrack provides a guided workflow for STM dynamics analysis:
- Load STM sequences from MPP movies or ordered STM frame series.
- Inspect metadata and navigate frames with playback-like controls.
- Preprocess difficult sequences using horizontal artifact repair and BM3D denoising.
- Register frame-to-frame drift using several registration backends.
- Create tracking seeds manually or from YOLO detections.
- Track particles or step edges across time using mask, point and edge-based backends.
- Review and correct tracks interactively in the GUI.
- Export measurements and sessions for reproducible downstream analysis.
The application stores particle and edge annotations in frame coordinates and provides aligned/expanded aligned views mainly for review and registration quality control.
Supported Input Data
NanoTrack currently targets grayscale STM-like image sequences.
| Input type | Use in NanoTrack |
|---|---|
.mpp | Native STM movie / multi-frame sequence loader. Frame timing metadata is parsed when available. |
.stp | Single STM frame; multiple .stp files can be loaded as an ordered frame series. |
.s94 | Single STM frame; multiple .s94 files can be loaded as an ordered frame series. |
For .stp and .s94 frame series, the frames must have matching image dimensions. The application loads them in the order selected by the user. A reverse-open option is available for sequences where the experimental order needs to be inverted.
Main Workflow
1. Load a Sequence
Use:
File -> Open STM...or, when needed:
File -> Open STM Reverse...NanoTrack creates an internal STMSequence with:
- raw frame stack,
- source path or source file list,
- frame count,
- image size and pixel scale,
- channel metadata,
- scan size,
- bias and setpoint metadata when present,
- frame timing when available from MPP metadata.
The main viewer shows the active frame and provides navigation through the sequence. Individual frames can be excluded from future analysis and exports.
2. Inspect and Preprocess
The central viewer is supported by metadata, frame navigation, overlay layers and side panels for tracking tools.
Available preprocessing tools include:
- horizontal dropout repair for line-like missing or corrupted scan artifacts,
- repair preview and apply-to-all workflow,
- BM3D denoising preview,
- BM3D batch application to all frames,
- optional viewing of cached preprocessed frames in the main viewer.
The preprocessing tools are intended to improve trackability and edge detection quality. They do not replace manual review.
3. Register the Sequence
NanoTrack includes a registration workflow for drift and frame alignment inspection.
Toolbar actions:
Registration Preview...
Run Registration...
View Registration...
Show Aligned
Show Expanded AlignedAvailable registration backends in the GUI:
- Phase correlation,
- Tile + RANSAC,
- ECC translation,
- Optical-flow median shift,
- Deep matcher translation.
Registration results include per-frame diagnostics such as:
dx,dyshift,- shift magnitude,
- quality score,
- status,
- phase peak ratio,
- ECC score,
- inlier tile count,
- tile residual,
- optical-flow MAD.
Registration outputs can be exported to CSV as metrics and summary files.
4. Create Particle Seeds
Particle tracking starts from seed bounding boxes.
NanoTrack supports two main seed sources.
Manual Seeds
Use the bbox tools panel to:
- set a default bbox size,
- place a bbox on the current frame,
- add it as a seed track,
- load a track bbox for correction,
- save a manual correction,
- resume a track from the corrected geometry.
YOLO Seed Detection
NanoTrack includes a YOLO seed-detection panel. It can:
- discover local YOLO model checkpoints,
- run detection on the current frame,
- run detection on all frames,
- scale detection boxes around their centers,
- edit selected detections,
- select or deselect detections globally or per-frame,
- convert selected detections into tracking seeds.
YOLO checkpoints are discovered from:
nanotrack/yolo_models/Supported checkpoint suffixes include:
.pt, .pth, .onnx, .engineYOLO inference requires ultralytics and a compatible torch installation.
5. Track Particles Over Time
NanoTrack separates the seed creation step from the tracking step. After seeds are created, the user can run tracking for a selected seed or for all seeds.
Current mask-tracking backends exposed by the UI:
- SAM2,
- DAM4SAM,
- SAMURAI.
The tracking result is stored as frame-indexed annotations for each particle track. Depending on the backend and review state, annotations may include:
- bbox geometry,
- segmentation mask,
- visibility state,
- source backend,
- source view,
- per-frame metrics.
Particle metrics include:
- area in pixels,
- area in
nm²when calibration is available, - perimeter in pixels,
- perimeter in
nmwhen calibration is available, - intensity sum,
- mean intensity,
- maximum intensity.
6. Create and Track Step-Edge ROIs
NanoTrack also supports step-edge workflows through polygon ROIs and edge polylines.
The polygon ROI panel allows the user to:
- draw a polygon ROI on a frame,
- preview edge detection inside the ROI,
- run edge detection over a selected frame range,
- load and correct the current edge,
- save manual edge corrections,
- resume edge tracking,
- re-detect one frame or a range,
- run hybrid edge stabilization using point trackers.
Supported edge-detector backends in the UI:
- DexiNed,
- TEED,
- NBED,
- DDN,
- PiDiNet,
- UAED,
- MuGE.
Available edge polyline extraction/refinement options include:
- graph path extraction,
- PCA + binning,
- combined refinement score,
- edge-probability refinement,
- gradient-based refinement,
- configurable normal-direction refinement radius.
Edge metrics include:
- polyline length in pixels,
- length in
nmwhen calibration is available, - RMS roughness,
- RMS roughness in
nmwhen calibration is available, - mean curvature,
- maximum curvature,
- waviness amplitude.
7. Hybrid Edge Stabilization with Point Trackers
For step-edge workflows, NanoTrack can sample control points from an anchor polyline and stabilize or propagate the edge over time using point-tracker backends.
Point-tracker backends currently wired through subprocess launchers:
- TAPIR,
- LocoTrack,
- Track-On-R.
These trackers are optional and require separate local installations, model checkpoints and backend configuration.
8. Review Results
NanoTrack provides dedicated result dialogs for:
- particle tracks,
- step-edge tracks,
- registration quality.
The result dialogs are intended for manual review before export. They expose tables and plots for checking whether tracks, frame exclusions, registration quality and computed metrics are suitable for downstream analysis.
9. Export and Save Sessions
NanoTrack supports reproducible session storage and CSV exports.
Session Format
Native session files use:
.nanotrackThe session file is a zip-backed project snapshot. It preserves:
- source sequence reference,
- active frame,
- excluded frames,
- particle tracks,
- edge tracks,
- YOLO detections,
- draft bboxes,
- draft polygon ROIs,
- draft edge polylines,
- repair and denoising cache metadata,
- registration results,
- aligned-view flags.
Particle Result Export
Particle exports produce:
<name>_metrics.csv
<name>_summary.csvThe metrics file contains per-track, per-frame measurements. The summary file aggregates per-track statistics.
Edge Result Export
Edge exports produce:
<name>_edge_metrics.csv
<name>_edge_summary.csvThe metrics file contains per-edge, per-frame geometry. The summary file aggregates per-edge statistics.
Registration Export
Registration exports produce:
<name>_registration_metrics.csv
<name>_registration_summary.csvThese files are useful for documenting drift correction quality and excluding problematic frames.
Typical Use Cases
Time-Resolved STM Particle Tracking
Use NanoTrack to follow bright particles, adsorbates, defects or local features across STM movies. The workflow combines manual or YOLO-based seeding with mask tracking and quantitative per-frame measurements.
STM Drift Inspection and Registration QA
Use registration preview and batch registration to inspect how much a sequence drifts over time. Export registration metrics to document shift, quality and failure modes.
Step-Edge Dynamics
Use polygon ROIs and edge-detection backends to identify step edges and track their position across a frame sequence. Edge metrics can quantify length, roughness, curvature and waviness.
Benchmarking Tracking Methods
NanoTrack is useful as a testbed for comparing classical registration, mask tracking, point tracking and edge-detection backends on the same STM sequence.
Manual Curation Before Quantitative Analysis
The application is not intended to be a fully automatic black-box pipeline. It supports manual seed placement, manual correction, track deletion, frame exclusion and result review.
Building Reproducible STM Movie Analyses
Use .nanotrack sessions to preserve the analysis state and CSV exports to move results into notebooks, statistical analysis tools or manuscript figures.
Installation and Launch
A minimal local workflow is:
python -m venv .venv
# Windows
.venv\Scripts\activate
# Linux/macOS
source .venv/bin/activate
pip install PyQt6 pyqtgraph numpy scipy scikit-image matplotlib
python -m nanotrack.mainDepending on the selected workflow, additional optional dependencies may be required:
pip install bm3d
pip install ultralytics torchExternal AI backends such as SAM2, DAM4SAM, SAMURAI, TAPIR, LocoTrack, Track-On-R, TEED, NBED, DDN, PiDiNet, UAED, MuGE and DexiNed may require separate repositories, checkpoints and dedicated Python or conda environments.
Optional Subprocess Backends
NanoTrack uses subprocess wrappers for several heavy external AI models. This makes it possible to keep the main PyQt application environment separate from tracker-specific environments.
Point trackers are configured through environment variables with prefixes such as:
NANOTRACK_TAPIR_PYTHON
NANOTRACK_TAPIR_REPO
NANOTRACK_TAPIR_CHECKPOINT
NANOTRACK_TAPIR_DEVICE
NANOTRACK_TAPIR_TIMEOUT_SEC
NANOTRACK_LOCOTRACK_PYTHON
NANOTRACK_LOCOTRACK_REPO
NANOTRACK_LOCOTRACK_CHECKPOINT
NANOTRACK_LOCOTRACK_DEVICE
NANOTRACK_LOCOTRACK_TIMEOUT_SEC
NANOTRACK_TRACKONR_PYTHON
NANOTRACK_TRACKONR_REPO
NANOTRACK_TRACKONR_CHECKPOINT
NANOTRACK_TRACKONR_DEVICE
NANOTRACK_TRACKONR_TIMEOUT_SECMask trackers and edge detectors currently also rely on backend configuration files and local model paths. In a clean deployment, these paths should be reviewed and replaced with workstation-specific paths or a more portable configuration layer.
Development Status
NanoTrack is an active development application. The core GUI, sequence loading, registration workflow, seed management, session persistence, particle metrics, edge metrics and CSV exports are present in the repository, but several AI-assisted tracking backends depend on optional external environments.
Current status summary:
- Stable enough for internal experimental workflows: sequence loading, manual seeds, session save/load, registration preview/results, particle and edge result exports.
- Backend-dependent: YOLO, SAM2, DAM4SAM, SAMURAI, TAPIR, LocoTrack, Track-On-R and deep edge detectors.
- Requires manual review: track quality, edge extraction quality, failed frames, frame exclusions and registration status.
- Not yet a turnkey public package: external model paths, checkpoints and environment setup may need project-specific configuration.
Feature Highlights
- STM movie and frame-series viewer,
.mpp,.stpand.s94support,- frame metadata and playback-style navigation,
- horizontal artifact repair and BM3D denoising,
- registration preview and batch registration,
- phase, Tile+RANSAC, ECC, optical-flow and deep-matcher registration options,
- manual bbox seeds and YOLO seed detection,
- SAM2 / DAM4SAM / SAMURAI mask tracking,
- TAPIR / LocoTrack / Track-On-R point tracking for edge stabilization,
- polygon ROI workflow for step-edge tracking,
- DexiNed / TEED / NBED / DDN / PiDiNet / UAED / MuGE edge backends,
- particle area, perimeter and intensity metrics,
- edge length, roughness, curvature and waviness metrics,
- registration QA metrics and CSV export,
.nanotracksession save/load.
License
The repository is distributed under the MIT License.