Software / 2026
Nanoparticle Analyzer (NaParA)
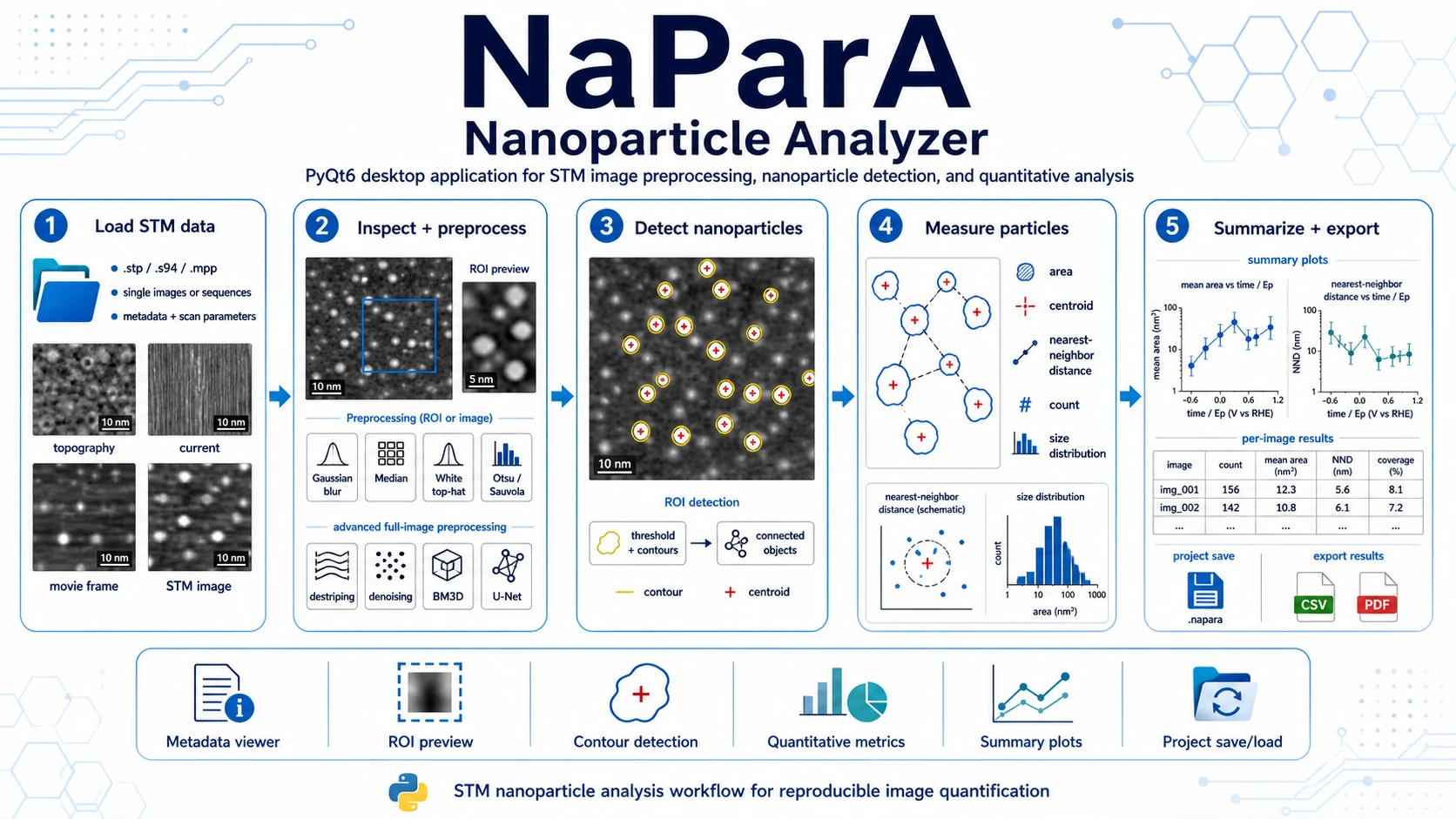
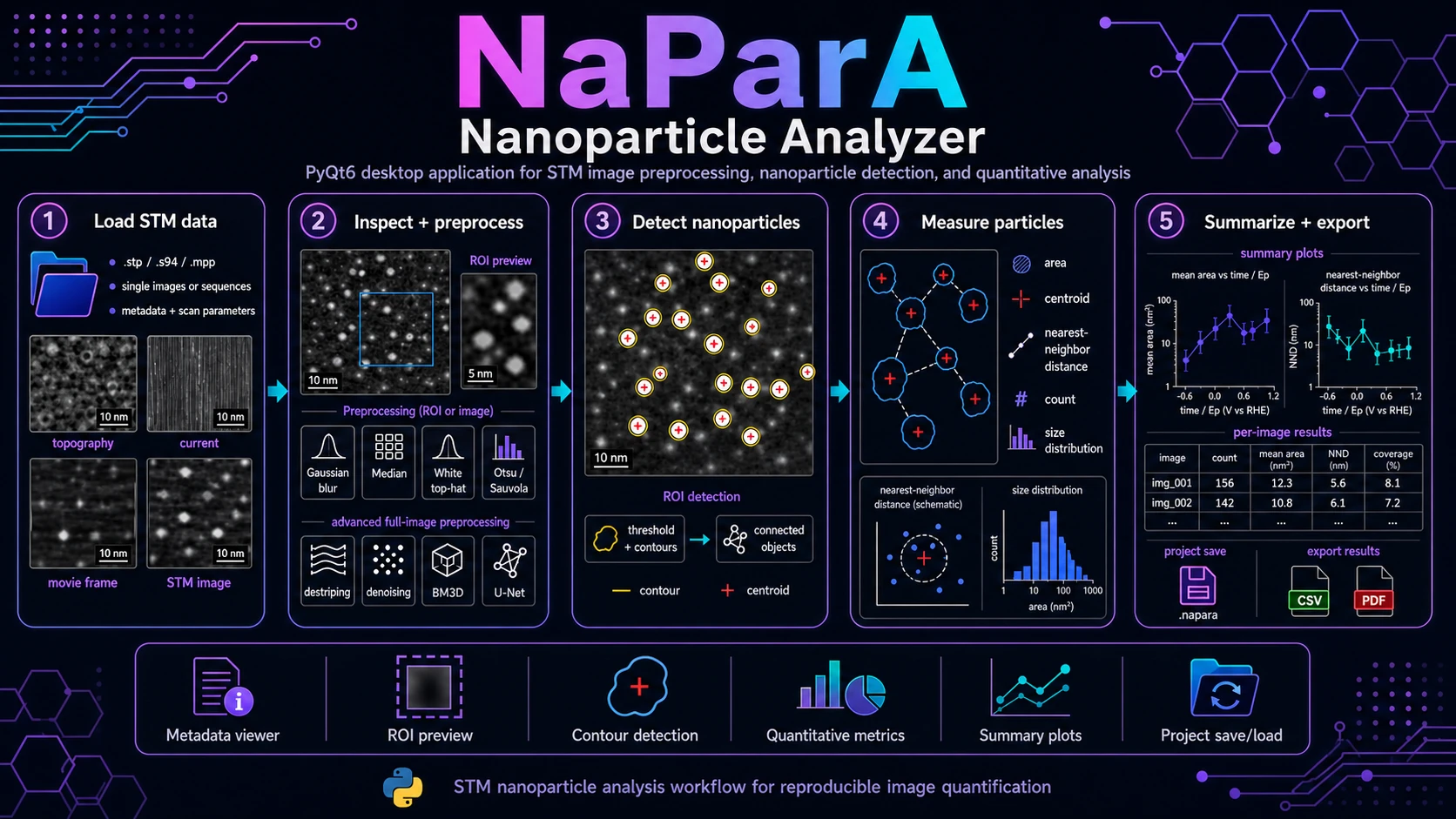
PyQt6 desktop application for STM image analysis: data loading, metadata inspection, ROI definition, preprocessing, nanoparticle-contour detection, geometric measurements and result export.
Overview
Nanoparticle Analyzer (NaParA) is a PyQt6 desktop application for STM image inspection, preprocessing, nanoparticle detection and quantitative image analysis. It is designed for research workflows where the user needs to load STM files, inspect metadata and calibration, define a region of interest, detect nanoparticle-like objects, measure their geometry and compare results across images or image sequences.
NaParA turns STM images into reproducible nanoparticle-level measurements: contours, centroids, areas, nearest-neighbour distances, per-image summaries, plots and CSV exports.
What NaParA Does
NaParA focuses on ROI-driven nanoparticle analysis in STM images. The application provides a practical workflow for datasets where nanoparticles, islands, bright features or similar nanoscale objects need to be detected and measured repeatedly across images.
Core functions include:
- loading STM data from supported experimental file formats,
- preserving calibration and metadata such as scan size, pixel scale, channel type, bias, current, scan angle and frame index,
- displaying STM images in physical units, with axes calibrated in nanometres when metadata are available,
- creating and editing a rectangular region of interest,
- previewing the selected ROI with the same display levels/LUT as the main image,
- applying lightweight ROI preprocessing before detection,
- running threshold-based contour/object detection inside the ROI,
- storing detected objects as editable contour overlays,
- computing per-object area, centroid and nearest-neighbour distance,
- visualizing area and nearest-neighbour distributions per image,
- plotting mean area and mean nearest-neighbour distance across an image series,
- exporting tabular results and plots for downstream analysis.
Supported Input Data
The current loader supports STM files from the repository readers:
| Format | Current behavior |
|---|---|
.stp | Omicron single-frame STM image. Header metadata and binary image payload are parsed. |
.s94 | SPECS STM image. Topography/current channels and physical scan metadata are parsed when available. |
.mpp | Omicron movie/sequence file. The file is expanded into individual STMImage frames. |
Loaded images are represented internally as STMImage objects containing the raw pixel array, dimensions, physical scan size, offsets, scan angle, bias, setpoint/current, image type and optional preprocessed data.
Main NaParA Workflow
1. Load STM Images
Start the application from the repository root:
python -m napara.mainIn the main window, add one or more STM files from the image list panel. NaParA accepts multiple .stp and .s94 images, and can expand .mpp files into frame sequences. Each loaded image appears in the image list and can be selected independently.
When an image is selected, the metadata panel shows the filename, pixel dimensions, physical scan size, pixel scale and channel type. The main viewer displays the image with calibrated axes when nm/px information is available.
2. Inspect the Image and Define an ROI
NaParA is built around ROI-based analysis. Use the ROI tools to create or reset a rectangular ROI on the active image. The ROI is stored per image and can be adjusted interactively.
The ROI preview updates from the active image and reflects the current processing settings. This makes it possible to tune detection parameters locally before committing detections to the image overlay.
3. Apply Preprocessing
NaParA has two levels of preprocessing.
The lightweight ROI/image pipeline in the main processing panel includes:
- Gaussian blur,
- median filtering,
- white top-hat filtering,
- Otsu or Sauvola thresholding,
- minimum object-area cleanup,
- contour/object detection toggle.
The full-image preprocessing dialog provides heavier operations for difficult STM data, including:
- median filtering,
- plane or polynomial leveling,
- scan-line correction and destriping,
- LOWESS line-trend removal,
- RANSAC line-baseline correction,
- Hough-guided streak removal,
- deconvolution,
- wavelet denoising,
- non-local means,
- Perona-Malik diffusion,
- directional total variation,
- BM3D,
- directional morphological reconstruction,
- optional U-Net denoising when compatible PyTorch weights are available.
Processed images can be used as the source for ROI preview and detection while preserving the original loaded data.
4. Detect Nanoparticles in the ROI
Enable thresholding and contour/object detection, then run detection for the active ROI. The detector applies the configured preprocessing, thresholding and connected-contour extraction inside the ROI.
Detected objects are stored as contour overlays. Each object receives an ID, a contour polygon in pixel coordinates, a centroid and an area. Existing detections inside the active ROI are replaced when a new detection pass is committed, while detections outside the ROI are preserved.
5. Review, Edit and Measure Detections
The detections dock lists the current image detections with:
- object ID,
- area in
nm²when calibration is available, - nearest-neighbour object ID,
- nearest-neighbour distance in
nm.
The user can show or hide contours, show or hide labels, delete selected detections, clear all detections for the active image and compute nearest neighbours. Nearest-neighbour distances are calculated from detected centroids in calibrated nanometre coordinates using a KD-tree.
6. Summarize and Export Results
The results dialog provides two analysis views:
- Per image - area distribution and nearest-neighbour distribution for the selected image, shown as KDE/histogram plots when enough data are available.
- Summary - mean area and mean nearest-neighbour distance across all loaded images.
The summary x-axis can use:
- image index,
- imported time values,
- imported electrochemical potential values,
Ep vs RHE.
Result export writes a folder named wyniki with:
summary.csv- one row per image,detections_long.csv- one row per detected object,summary_mean_area.png,summary_mean_nn.png,- per-image area-distribution PNG files,
- per-image nearest-neighbour PNG files.
7. Save and Reload Projects
NaParA projects are stored as .napara files. The project archive preserves:
- the image list and relative source paths,
- active image index,
- ROI rectangles,
- detections,
- nearest-neighbour values,
- optional preprocessed arrays.
The application also supports quick save/load slots under a local saves/ directory.
Typical Use Cases
Nanoparticle Population Analysis
Use NaParA to count particles and compare particle-size distributions across STM images. This is useful when the goal is to quantify how particle coverage, apparent size or nearest-neighbour spacing changes between samples, preparation conditions or electrochemical states.
Potential- or Time-Dependent STM Series
For image series where the filename or imported metadata can be matched to time or electrode potential, the results dialog can plot mean particle area and mean nearest-neighbour distance versus image index, time or Ep vs RHE.
ROI-Specific Detection on Heterogeneous Surfaces
Many STM images contain regions with different contrast, drift, streaks or background. NaParA allows detection to be restricted to a user-defined ROI, so parameters can be tuned for the region that is scientifically relevant instead of forcing one global threshold over the entire frame.
Preprocessing and Denoising Experiments
The full-image preprocessing dialog is useful for testing how leveling, destriping, denoising, deconvolution and morphological operations affect downstream object detection. This helps compare processing strategies without separating them from quantitative outputs.
Dataset Curation for Later Workflows
NaParA stores detected contours and measurements in a project file and exports long-form CSV results. This makes it suitable for building curated reference datasets and for comparing detection strategies while the software is still evolving.
Installation and Launch
The current archive does not include a pinned requirements.txt, so dependencies should be installed according to the workflows that will be used.
A practical base environment for the current NaParA GUI is:
pip install PyQt6 pyqtgraph numpy scipy scikit-image matplotlib statsmodels bm3d torchbm3d and torch are listed in the practical base environment because the current development code imports preprocessing/U-Net helper modules through the GUI path. If those imports are later made lazy, torch can become optional for users who do not run the U-Net denoiser.
For HDF5 pair export, also install:
pip install h5pyCurrent Development Status
NaParA is a research-oriented desktop application in active development. The current codebase already contains a working path for loading STM images, ROI preview, preprocessing, contour detection, per-object metrics, nearest-neighbour analysis, result plots, CSV export and .napara project persistence.
Important current-state notes:
- the primary NaParA detection path is ROI-driven and threshold/contour based,
- PDF/report export is not implemented in the current NaParA code; current result export is CSV plus PNG plots,
- U-Net denoising requires compatible PyTorch weights and model weights; the code raises a clear error when weights are missing,
- for publication-grade use, every dataset should still be visually reviewed after preprocessing and detection.
NaParA Scope
NaParA is focused on object-level STM image quantification: nanoparticle-like features, contours, centroids, areas, nearest-neighbour distances, image-series summaries and reproducible export.
License
The repository is distributed under the MIT License.